load xlr8r
![]()

define a model
It might help to open the xCellerator palette (from the Palettes menu, above)
This is a model of a simple enzymatic transition that is initiated by an input signal “input” and the product is degraded at a constant rate d.

![]()
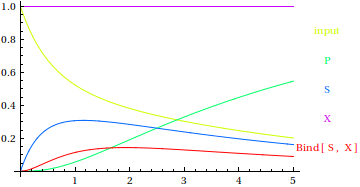
Test the model by running a simulation.
All parameters that are not set will default to a value of 1. In the following simulation, the parameter d=0 but all other parameters such as k1,k2, k3,k4 are automatically set to 1. This is useful for testing a model, but eventually you will have to tune your parameters to the correct values.
All initial conditions (values of variables such as S, P, X, etc at t=0) will default to zero. In the following simulation, the initical value of “input” and “X” are 1, and the initial values of S and P are both zero, since they are not specified.
When running your own simulations you should take special care here to make sure that the units of all of your parameters and initial conditions are consistent, e.g., if you mix nanoMoles with microMoles, make sure to apply to appropriate conversion factors to your values.
We use the frozen option to keep the enzyme X fixed. This can also be done by adding a reaction {Ø⇔X,kf,kr} with kf/kr set to the desired steady state value. However, if we use the reaction, we will have to wait for the reaction to reach steady state; the frozen option puts it there immediately.


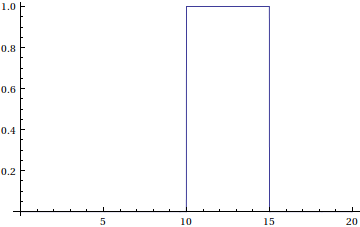
Instead of specifying an input for the reactang S, we want to keep it zero until such time that an input signal turns it on. We define a stimulus function:
![]()
![]()
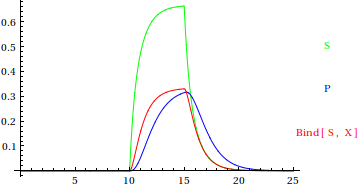
run a simulation using the stimulus function for the variable “input”
use runPlot instead of automatic plotting to give more control over plots.

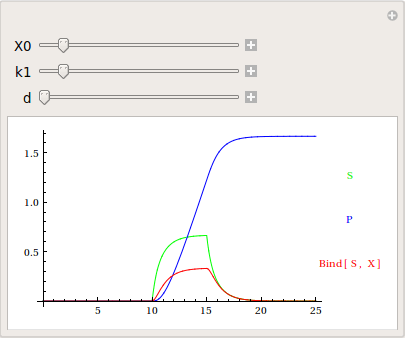
Use manipulate to play with some of the parameters and initial conditions.